This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
What is phylogenetics?
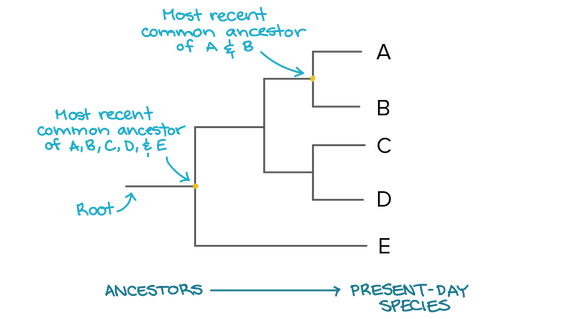
Phylogenetics is the study of evolutionary relationships among biological entities. Phylogenetics helps us understand how species and genes evolve. Evolutionary relationships can be represented by a phylogenetic tree. On the tree, each branch represents an individual species or gene. The node connecting branches represents the most recent common ancestors the species shared before diverging. [1]
Figure 1: A phylogenetic tree diagram
Types of Phylogenetic Trees [2]
Neighbor Joining:Uses similarity scores from percent identity to determine which species are most closely related to each other. Branch length is calculated from these scores.
Maximum Likelihood: This method builds a tree using another method and then adjust branch lengths based on maximum likelihood.
Average Distance: Uses similarity scores between species to determine which species are most closely related and uses equal branch length from a common ancestor.
Maximum Likelihood: This method builds a tree using another method and then adjust branch lengths based on maximum likelihood.
Average Distance: Uses similarity scores between species to determine which species are most closely related and uses equal branch length from a common ancestor.
MSH6 Phylogenetic Tree Construction
1. Protein sequences of homologs to MSH6 from various organisms are found utilizing Ensemble. The sequences are annotated in a FASTA format.
2. The sequences are aligned using ClustalOmega.
3. A tree is constructed using MEGA.
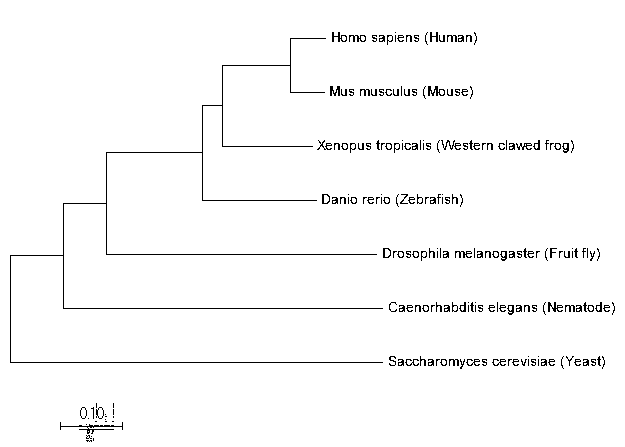
Neighbor Joining Tree for MSH6
Conclusions
The evolutionary relationships between the MSH6 protein homologs are shown through the construction of a phylogenetic tree. BLAST contained the percent similarity between these homologs, and the phylogenetic tree confirms the strength of these relationships. For example, humans and mice have a high similarity between their MSH6 proteins, and they are the most closely related, while yeast has a much lower similarity and is less closely related. The different methods of phylogenetic tree construction yielded the same resulting tree suggesting a high confidence in these evolutionary relationships between these MSH6 protein homologs.
References:
[1] EMBL-EBI Train Online. (2019). What is phylogenetics?. Retrieved from https://www.ebi.ac.uk/training/online/course/introduction-phylogenetics/what-phylogenetics.
[2] Yang, Z and Rannala, B. (2012). Molecular phylogenetics: principles and practices. Nat Rev Genet. 13(5): 303-314. doi: 10.1038/nrg3186
Images:
Header: https://images.slideplayer.com/25/8111838/slides/slide_1.jpg
Figure 1: cdn.kastatic.org/ka-perseus-images/aa95c701ebf845d93fed8362da63cbcc8439fb31.png
[1] EMBL-EBI Train Online. (2019). What is phylogenetics?. Retrieved from https://www.ebi.ac.uk/training/online/course/introduction-phylogenetics/what-phylogenetics.
[2] Yang, Z and Rannala, B. (2012). Molecular phylogenetics: principles and practices. Nat Rev Genet. 13(5): 303-314. doi: 10.1038/nrg3186
Images:
Header: https://images.slideplayer.com/25/8111838/slides/slide_1.jpg
Figure 1: cdn.kastatic.org/ka-perseus-images/aa95c701ebf845d93fed8362da63cbcc8439fb31.png